We take part in Kaggle/MICCAI 2020 challenge to classify Prostate cancer “Prostate cANcer graDe Assessment (PANDA) Challenge Prostate cancer diagnosis using the Gleason grading system”
From the organizer website:
With more than 1 million new diagnoses reported every year, prostate cancer (PCa) is the second most common cancer among males worldwide that results in more than 350,000 deaths annually. The key to decreasing mortality is developing more precise diagnostics. Diagnosis of PCa is based on the grading of prostate tissue biopsies. These tissue samples are examined by a pathologist and scored according to the Gleason grading system. In this challenge, you will develop models for detecting PCa on images of prostate tissue samples, and estimate severity of the disease using the most extensive multi-center dataset on Gleason grading yet available.
The grading process consists of finding and classifying cancer tissue into so-called Gleason patterns (3, 4, or 5) based on the architectural growth patterns of the tumor (Fig. 1). After the biopsy is assigned a Gleason score, it is converted into an ISUP grade on a 1-5 scale. The Gleason grading system is the most important prognostic marker for PCa, and the ISUP grade has a crucial role when deciding how a patient should be treated. There is both a risk of missing cancers and a large risk of overgrading resulting in unnecessary treatment. However, the system suffers from significant inter-observer variability between pathologists, limiting its usefulness for individual patients. This variability in ratings could lead to unnecessary treatment, or worse, missing a severe diagnosis.
Automated deep learning systems have shown some promise in accurately grading PCa. Recent research, including two studies independently conducted by the groups hosting this challenge, have shown that these systems can achieve pathologist-level performance. However, these systems/results were not tested with multi-center datasets at scale.
Your work here will improve on these efforts using the most extensive multi-center dataset on Gleason grading yet. The training set consists of around 11,000 whole-slide images of digitized H&E-stained biopsies originating from two centers. This is the largest public whole-slide image dataset available, roughly 8 times the size of the CAMELYON17 challenge, one of the largest digital pathology datasets and best known challenges in the field. Furthermore, in contrast to previous challenges, we are making full diagnostic biopsy images available. Using a sizable multi-center test set, graded by expert uro-pathologists, we will evaluate challenge submissions on their applicability to improve this critical diagnostic function.
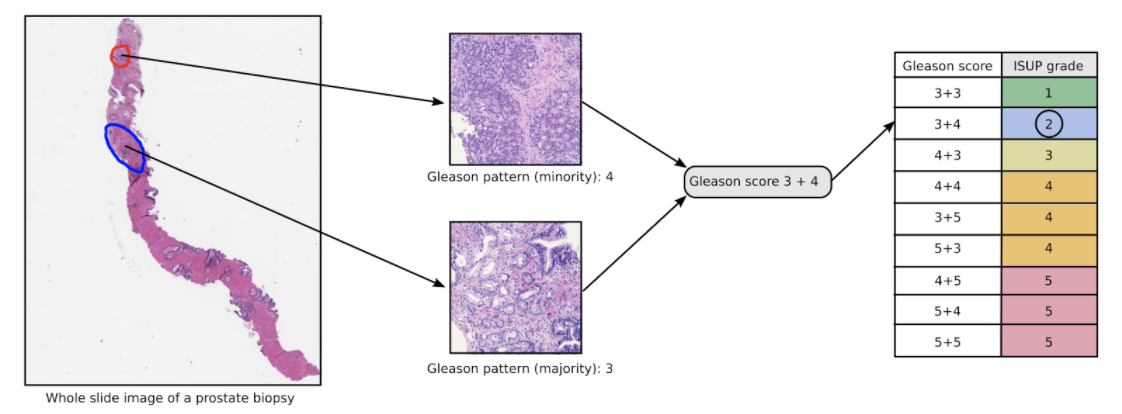
Figure 1: An illustration of the Gleason grading process for an example biopsy containing prostate cancer. The most common (blue outline, Gleason pattern 3) and second most common (red outline, Gleason pattern 4) cancer growth patterns present in the biopsy dictate the Gleason score (3+4 for this biopsy), which in turn is converted into an ISUP grade (2 for this biopsy) following guidelines of the International Society of Urological Pathology. Biopsies not containing cancer are represented by an ISUP grade of 0 in this challenge.
Radboud University Medical Center and Karolinska Institute have teamed up to organize this competition in collaboration with colleagues from Tampere University. The Computational Pathology Group (CPG) of the Radboud University Medical Center is a research group that develops computer algorithms to aid clinicians. Karolinska Institute’s Department of Medical Epidemiology and Biostatistics (MEB) includes an interdisciplinary research group to improve the diagnostics and treatment of prostate cancer. Together, they hope to further their existing research to make a significant impact on the healthcare of prostate cancer patients.
Challenge organizer team: Wouter Bulten, Geert Litjens, Hans Pinckaers, Peter Ström, Martin Eklund, Lars Egevad, Henrik Grönberg, Kimmo Kartasalo, Pekka Ruusuvuori, Tomi Häkkinen, Sohier Dane, Maggie Demkin.
Sponsors
The PANDA workshop at MICCAI 2020 is sponsored by ContextVision, Ibex and Google.



This competition is part of MICCAI 2020: the 23rd International Conference on Medical Image Computing and Computer Assisted Intervention. This year, MICCAI will be held from October 4th to 8th, 2020 in Lima, Peru. As part of the conference, the competition’s organizers will host a workshop on the results of the PANDA challenge.

more info: https://www.kaggle.com/c/prostate-cancer-grade-assessment


